cKBET: assessing goodness of batch effect correction for single-cell RNA-seq
Por um escritor misterioso
Last updated 04 abril 2025

lt;p>Single-cell RNA sequencing reveals the gene structure and gene expression status of a single cell, which can reflect the heterogeneity between cells. However, batch effects caused by non-biological factors may hinder data integration and downstream analysis. Although the batch effect can be evaluated by visualizing the data, which actually is subjective and inaccurate. In this work, we propose a quantitative method cKBET, which considers the batch and cell type information simultaneously. The cKBET method accesses batch effects by comparing the global and local fraction of cells of different batches in different cell types. We verify the performance of our cKBET method on simulated and real biological data sets. The experimental results show that our cKBET method is superior to existing methods in most cases. In general, our cKBET method can detect batch effect with either balanced or unbalanced cell types, and thus evaluate batch correction methods.</p>
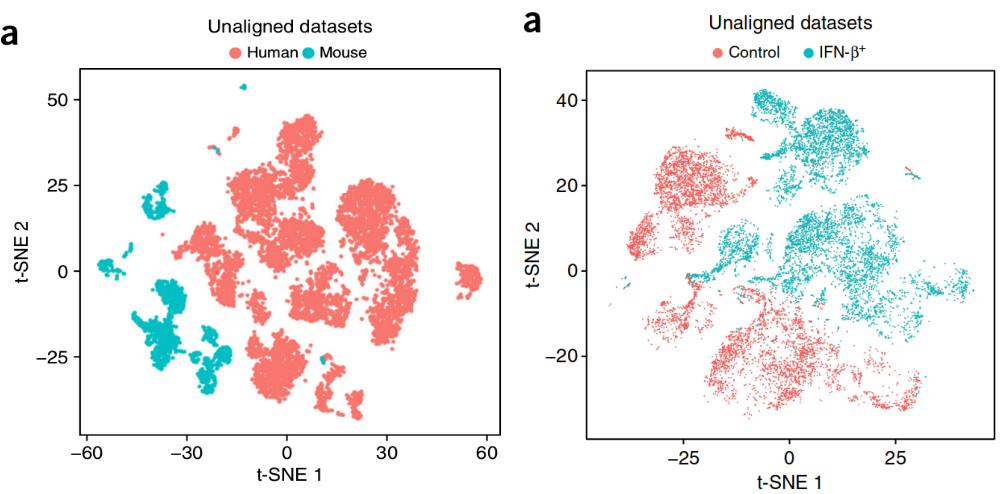
How to Batch Correct Single Cell. Comparing batch correction methods for…, by Nikolay Oskolkov
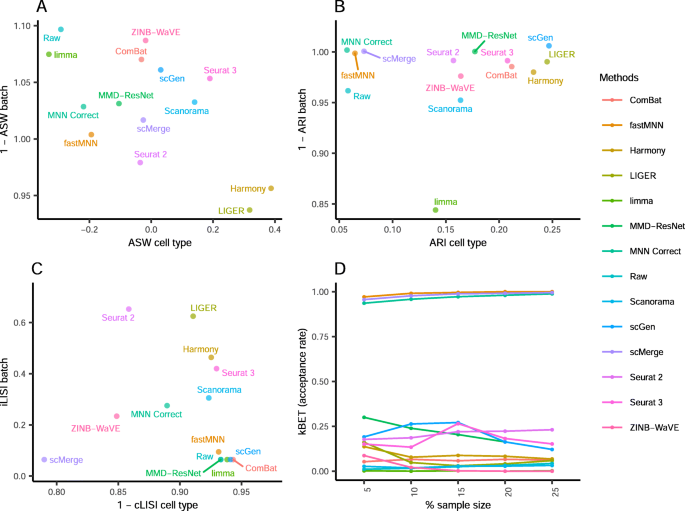
A benchmark of batch-effect correction methods for single-cell RNA sequencing data, Genome Biology
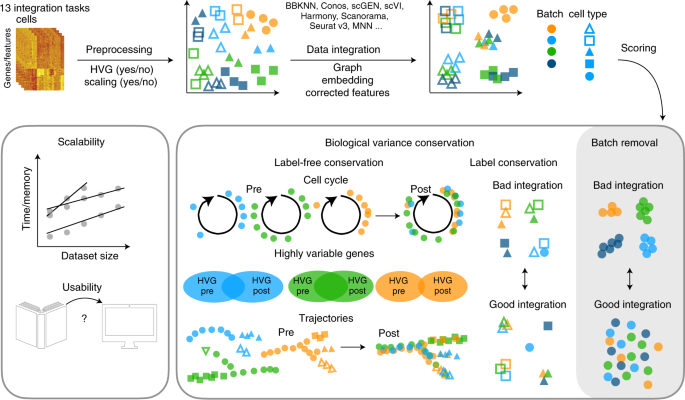
Benchmarking atlas-level data integration in single-cell genomics

Batch effects in single-cell RNA-sequencing data are corrected by matching mutual nearest neighbors
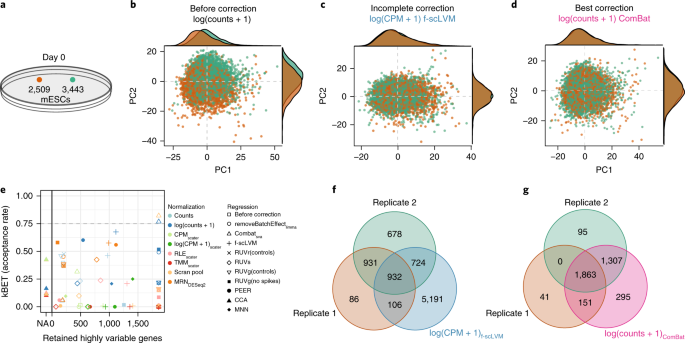
A test metric for assessing single-cell RNA-seq batch correction
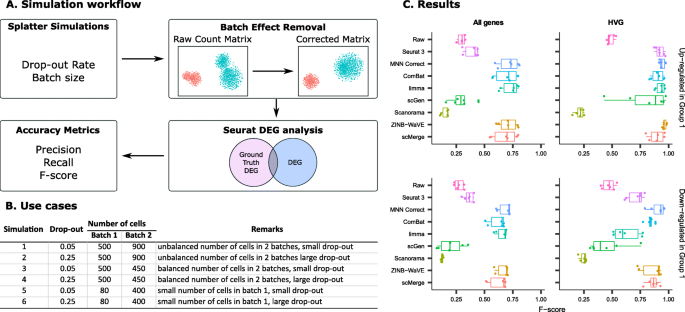
A benchmark of batch-effect correction methods for single-cell RNA sequencing data, Genome Biology

PDF) BERMUDA: A novel deep transfer learning method for single-cell RNA sequencing batch correction reveals hidden high-resolution cellular subtypes
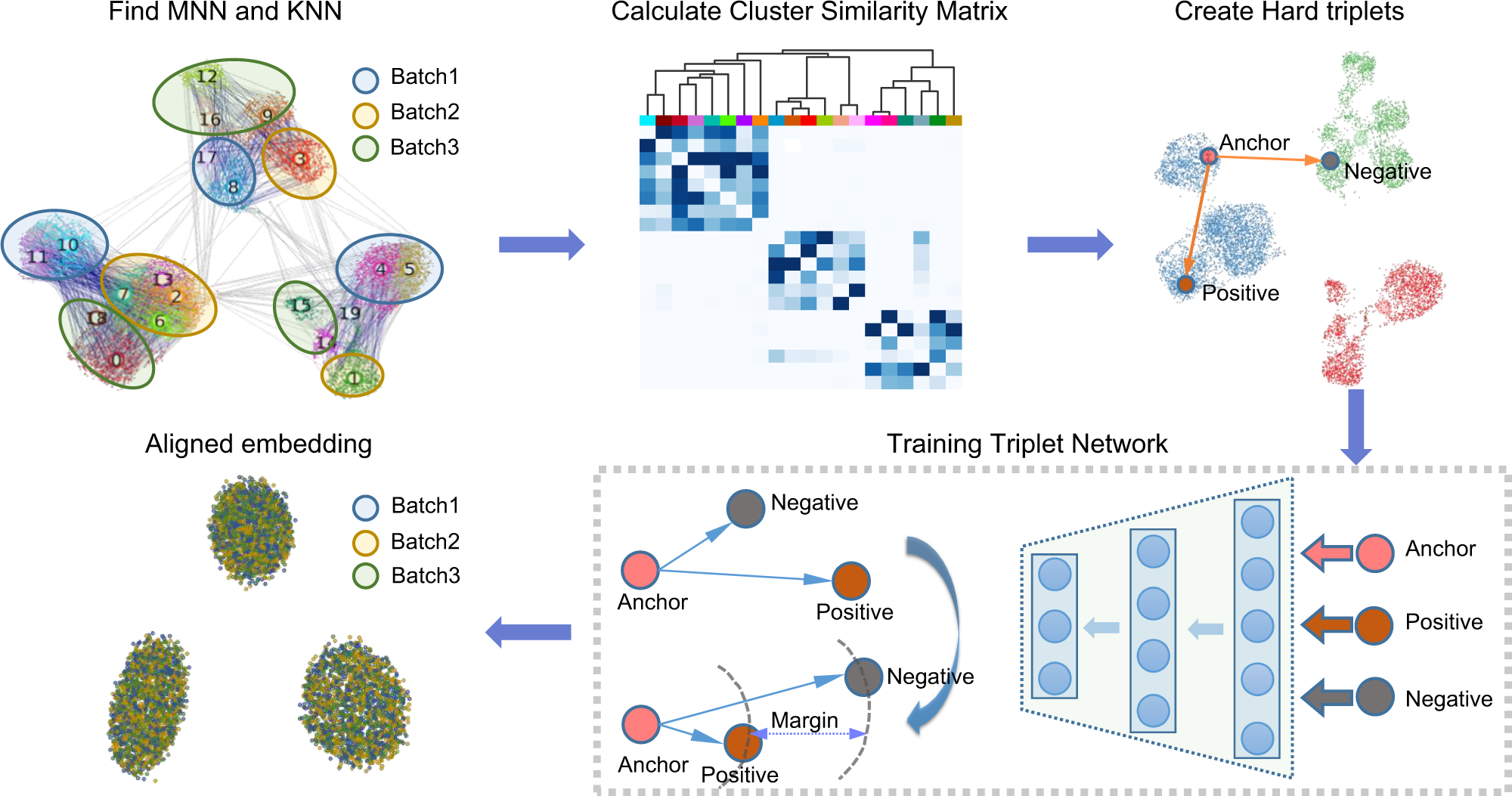
Batch alignment of single-cell transcriptomics data using deep metric learning

Assessment of batch-correction methods for scRNA-seq data with a new test metric

PDF) BEENE: Deep Learning based Nonlinear Embedding Improves Batch Effect Estimation
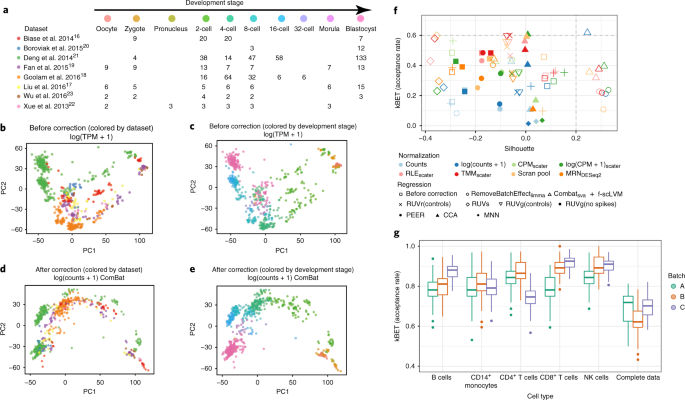
A test metric for assessing single-cell RNA-seq batch correction
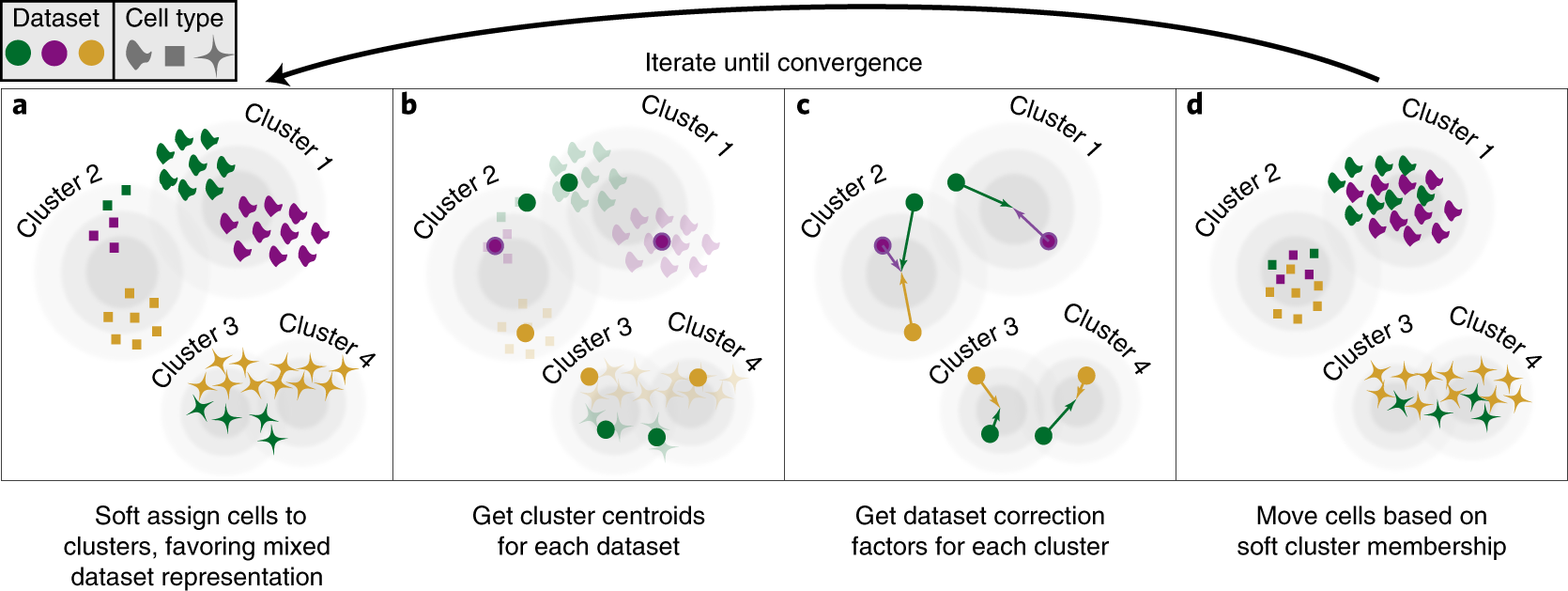
Fast, sensitive and accurate integration of single-cell data with Harmony
Recomendado para você
-
 OKbet Online Games: Enjoy - Issuu04 abril 2025
OKbet Online Games: Enjoy - Issuu04 abril 2025 -
 ckbetcam (ckbet) - Replit04 abril 2025
ckbetcam (ckbet) - Replit04 abril 2025 -
 Odd_Limit_6518 (u/Odd_Limit_6518) - Reddit04 abril 2025
Odd_Limit_6518 (u/Odd_Limit_6518) - Reddit04 abril 2025 -
 ckbet 🎱 【006.COM】 ckbet Revolucionando as Apostas Online com Recursos Inigualáveis #104 abril 2025
ckbet 🎱 【006.COM】 ckbet Revolucionando as Apostas Online com Recursos Inigualáveis #104 abril 2025 -
 ckbet plataforma04 abril 2025
ckbet plataforma04 abril 2025 -
 Browse thousands of 168jogo Com Sacar Ckbet Com95429 images for design inspiration04 abril 2025
Browse thousands of 168jogo Com Sacar Ckbet Com95429 images for design inspiration04 abril 2025 -
Download Ckbet on PC (Emulator) - LDPlayer04 abril 2025
-
 Ckbet – Oferecendo Uma Experiência de Apostas Segura e Conveniente Para Usuários Brasileiros – Portal G3704 abril 2025
Ckbet – Oferecendo Uma Experiência de Apostas Segura e Conveniente Para Usuários Brasileiros – Portal G3704 abril 2025 -
 CKBET Paga Mesmo? Dá pra ganhar dinheiro?04 abril 2025
CKBET Paga Mesmo? Dá pra ganhar dinheiro?04 abril 2025 -
 Spin, Bet, Win: Experience the Magic of CKBet Online Casino! - UrbanMatter04 abril 2025
Spin, Bet, Win: Experience the Magic of CKBet Online Casino! - UrbanMatter04 abril 2025
você pode gostar
-
 Como deixar a batata frita crocante: confira as DICAS DEFINITVAS para não errar04 abril 2025
Como deixar a batata frita crocante: confira as DICAS DEFINITVAS para não errar04 abril 2025 -
 CLASSROOM OF THE ELITE 3 TEMPORADA! QUANDO CHEGA NA CRUNCHYROLL?04 abril 2025
CLASSROOM OF THE ELITE 3 TEMPORADA! QUANDO CHEGA NA CRUNCHYROLL?04 abril 2025 -
 Techno House : r/Technoblade04 abril 2025
Techno House : r/Technoblade04 abril 2025 -
 ArtStation - Cute Dog GIF Set (Commission)04 abril 2025
ArtStation - Cute Dog GIF Set (Commission)04 abril 2025 -
 Mesa de ping pong mdp 15mm 1007 klopf c/ rodas, suporte E rede04 abril 2025
Mesa de ping pong mdp 15mm 1007 klopf c/ rodas, suporte E rede04 abril 2025 -
 CharlieIntel on X: Per new Call of Duty ads, the PlayStation 5 Slim Disc Edition - Call of Duty: Modern Warfare III console bundle will be $499. This is basically a new04 abril 2025
CharlieIntel on X: Per new Call of Duty ads, the PlayStation 5 Slim Disc Edition - Call of Duty: Modern Warfare III console bundle will be $499. This is basically a new04 abril 2025 -
 Matador de Demônios Tanjiro para colorir04 abril 2025
Matador de Demônios Tanjiro para colorir04 abril 2025 -
A sua vez vai chegar! ✍🏼 #tiktok #fypシ #viral #mckevin #motivacao #st04 abril 2025
-
 APRENDER JAPONÉS】¿Cómo se usa la palabra ”やばい Yabai”?04 abril 2025
APRENDER JAPONÉS】¿Cómo se usa la palabra ”やばい Yabai”?04 abril 2025 -
 Olimpia Scores, Stats and Highlights - ESPN04 abril 2025
Olimpia Scores, Stats and Highlights - ESPN04 abril 2025
